Overview
Model structure
The MEmilio library uses a modular organization of models where compartmental or aggregated models based on ODEs (ordinary differential equations) without and with Linear Chain Trick, IDEs (integro-differential equations), and SDEs (stochastic differential equations) share a maximum properties and interfaces (implemented in the memilio folder) to allow simple and straightforward model adaption with, e.g., demographic or spatial stratification (e.g. found in the memilio/mobility folder to create metapopulation models) as shown in the following figure. Agent-based models are furthermore harmonized with most structures such as parameters, contact patterns, and non-pharmaceutical interventions (e.g. found in memilio/epidemiology).
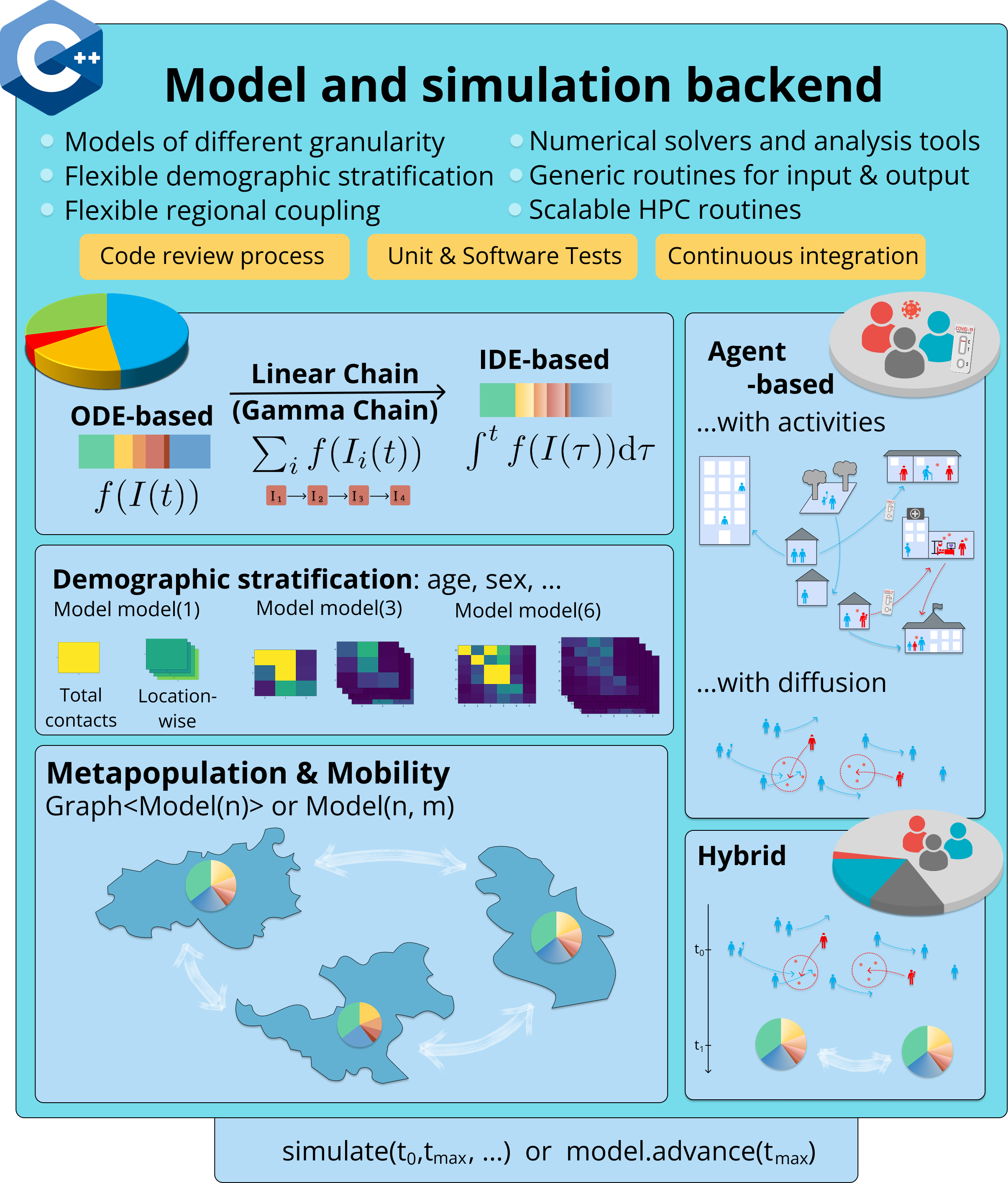
For a quick run through MEmilio’s functionality see Build instructions.
The MEmilio C++ project is organized as follows:
Main directory structure
The main directory structure in the cpp directory includes:
memilio/: Contains the core framework for developing epidemiological models
ad/: Algorithmic differentiation framework
compartments/: Classes for compartment models and simulations
data/: Data analysis functions
epidemiology/: Base classes for epidemiological modeling
geography/: Geographic region data (e.g. school vacations) and functions
io/: Input/output utilities for various formats (JSON, HDF5…)
math/: Mathematical utilities such as integrators (Euler, RK)
mobility/: Different Metapopulation mobility approaches
utils/: General helper functions (logging, etc.)
models/: Concrete implementation of epidemiological models
simulations/: Applications for scientific publications
examples/: Example applications demonstrating the use of the framework
tests/: Unit tests for framework and models
thirdparty/: Configuration of external dependencies
benchmarks/: Analyzing runtime performance
Build system
The project uses CMake as a build system with various configuration options. For an explanation, refer to Build instructions.