MEmilio - a high performance Modular EpideMIcs simuLatIOn software
Welcome
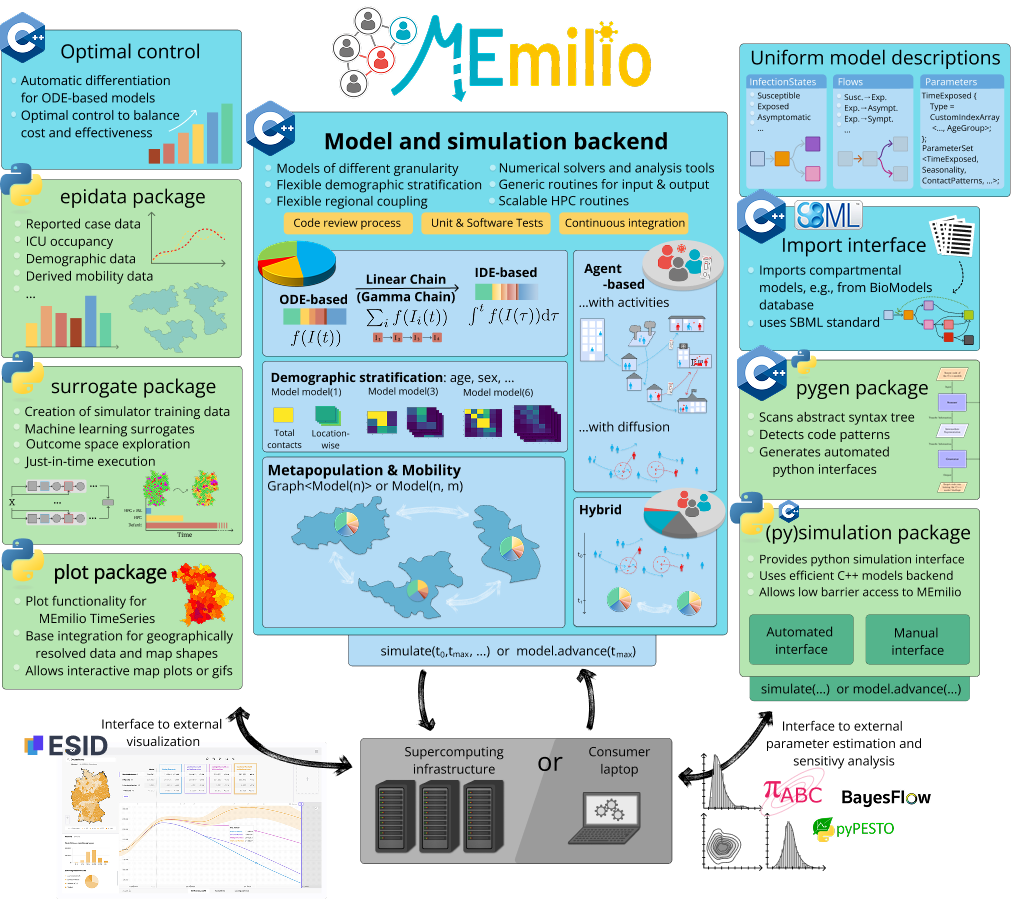
MEmilio implements various models for infectious disease dynamics, ranging from simple compartmental models to complex Integro-Differential and agent-based models. Its modular design enables the combination of different models with distinct mobility patterns. Through efficient implementation and parallelization in C++ and an easy-to-use python interface, MEmilio delivers cutting-edge and compute-intensive epidemiological models to broad range of applications and users, providing precise and high-resolution spatiotemporal infectious disease dynamics. MEmilio is continuously extended and is available open-source for community use.
If you use MEmilio, please cite our work.
Note
This framework is under active development, and, thus, obtains constantly new features or models. If you encounter a feature not yet documented, please contact us directly.